Mutations in the survival motor neuron (SMN1) gene cause spinal muscular atrophy (SMA), an autosomal recessive disorder characterised by degeneration of spinal cord motor neurons leading to progressive muscular weakness. SMN1 encodes an RNA-binding protein, SMN, which is complexed with Gemin proteins. Vertebrate Gemin3 is the only RNA helicase in the SMN complex. In Drosophila melanogaster, CG6539 was identified as the orthologue of vertebrate gemin3. It was shown to physically interact with SMN, and its loss results into larval/prepupal lethality. Before death, gemin3 mutant larvae exhibit declined mobility and expanded neuromuscular junctions.
Cauchi RJ, Davies KE, Liu J-L (2008) A Motor Function for the DEAD-Box RNA Helicase, Gemin3, in Drosophila. PLoS Genet 4(11): e1000265. doi:10.1371/journal.pgen.1000265
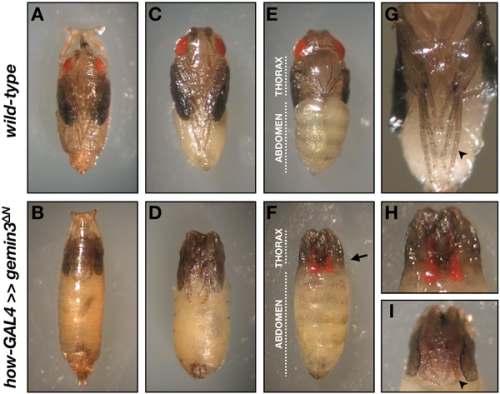
doi:10.1371/journal.pgen.1000265.g008
x

